S. Erattakulangara, K. Kelat, K. Burnham, R. Balbi, S.E. Gerard, D. Meyer, S.G. Lingala
"Open-source manually annotated vocal tract database for automatic segmentation from 3D MRI using Deep Learning: Benchmarking 2D and 3D convolutional and transformer networks"
Journal of Voice (in-press).
Data-download [Fig-share link]
Abstract of the article:
Objectives: Accurate segmentation of the vocal tract from MRI data is essential for various voice, speech, and singing applications. Manual segmentation is time-intensive and susceptible to errors. This study aimed to evaluate the efficacy of deep learning algorithms for automatic vocal tract segmentation from 3D MRI.
Study Design: This study employed a comparative design, evaluating four deep learning architectures for vocal tract segmentation using an open-source dataset of 3D MRI scans of French speakers.
Methods: Fifty-three vocal tract volumes from 10 French speakers were manually annotated by an expert vocologist, assisted by two graduate students in voice science. These included 21 unique French phonemes and 3 unique voiceless tasks. Four state- of-the-art deep learning segmentation algorithms were evaluated: 2D slice-by-slice U- Net, 3D U-Net, 3D U-Net with transfer learning (pre-trained on lung CT), and 3D transformer U-Net (3D U-NetR). The STAPLE algorithm, which combines segmentations from multiple annotators to generate a probabilistic estimate of the true segmentation, was used to create reference segmentations for evaluation. Model performance was assessed using the Dice coefficient, Hausdorff distance, and structural similarity index measure.
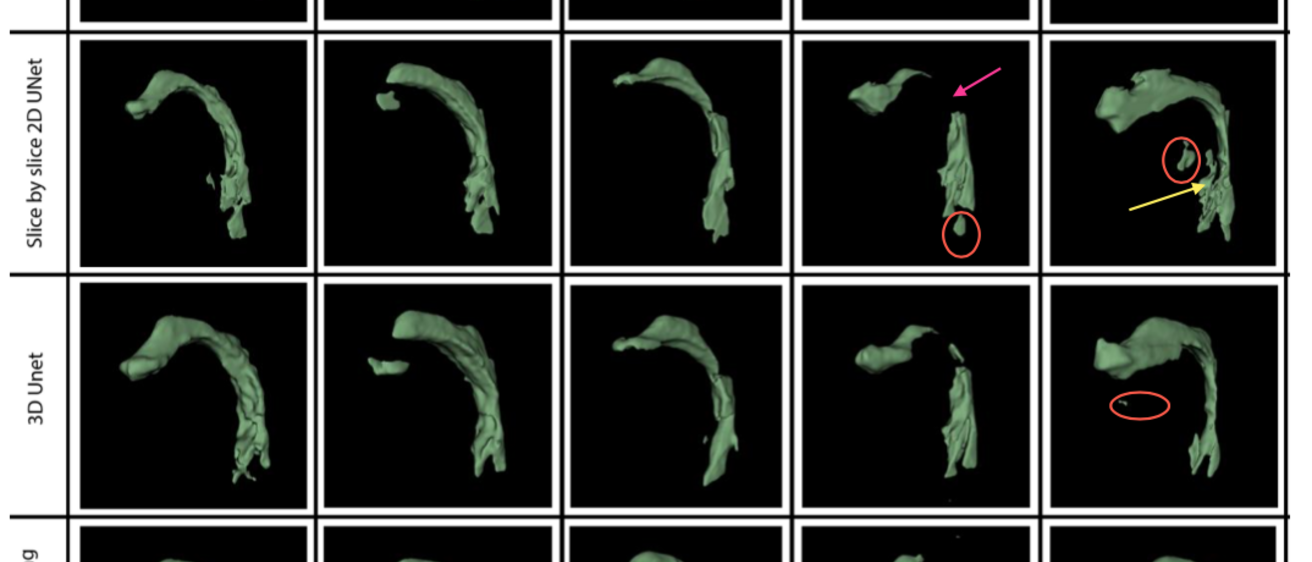
Results: The 3D U-Net and 3D U-Net with transfer learning models achieved the highest Dice coefficients (0.896 ± 0.05 and 0.896 ± 0.04, respectively). The 3D U-Net with transfer learning performed comparably to the 3D U-Net while using less than half the training data. It, along with the 2D slice-by-slice U-Net models demonstrated lower variability in HD distance compared to the 3D U-Net and 3D U-NetR models. All models exhibited challenges in segmenting certain sounds, particularly /kõn/. Qualitative assessment by a voice expert revealed anatomically correct segmentations in the oropharyngeal and laryngopharyngeal spaces for all models except the 2D slice- by-slice UNET, and frequent errors with all models near bony regions (eg. teeth).
Conclusions: This study emphasizes the effectiveness of 3D convolutional networks, especially with transfer learning, for automatic vocal tract segmentation from 3D MRI. Future research should focus on improving the segmentation of challenging vocal tract configurations and refining boundary delineations.